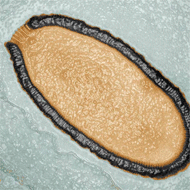
Scientists discover a new type of virus in frozen ground
French scientists have discovered an ancient virus that has survived for more than 30,000 years under frozen ground in north-eastern Siberia.
Researchers from the National Centre of Scientific Research (CNRS) say the virus - known as Pithovirus sibericum - poses no threat to animals or humans.
It belongs to a class of giant viruses - the only viruses that can be seen under optical microscopy. The discovery brings the number of known giant viruses up to three.
Scientists say their research shows viruses can survive in the permanently frozen layer of soil found in Arctic regions (known as permafrost) over geological time periods. According to the research team, this could have important public health implications.
Work must be done, they say, to provide a realistic estimate of the likelihood of viruses re-emerging after they were thought to be eradicated. Scientists are now working on a metagenomic study of permafrost.
Giant viruses infect amoeba such as Acanthamoeba and contain very large numbers of genes compared to common viruses such as AIDS or influenza.
According to research published in Proceedings of the National of Sciences this week, Pithovirus is reminiscent of another giant virus, Pandoravirus, but in fact they are very different.
Pithovirus contains around 500 genes, far fewer than the 2,500 Pandoravirus can carry. In addition, the new virus is made up of hundreds of proteins, whereas Pandoravirus contains only one or two.
The research team discovered the 30,000-year-old virus had almost nothing in common with other giant viruses and belongs to a new family.
Image © Julia Bartoli & Chantal Abergel, IGS, CNRS/AMU



 Two independent vets have launched a podcast to help owners strengthen their bond with pets. Dr Maggie Roberts and Dr Vanessa Howie, who have worked in both veterinary practice and major charities, are keen to use their experience to enable people to give pets a better life.
Two independent vets have launched a podcast to help owners strengthen their bond with pets. Dr Maggie Roberts and Dr Vanessa Howie, who have worked in both veterinary practice and major charities, are keen to use their experience to enable people to give pets a better life.